In semiconductor manufacturing, maintaining consistent process performance across wafers and across batches is critical for device performance. One of the most powerful tools for monitoring this consistency is the Statistical Process Control (SPC) Chart. In this tutorial, we walk through how to apply SPC charts in R to simulate thin film deposition data, covering X-bar and R charts, control limit calculations, and practical interpretation for process engineers.
If you are new to R for data analysis, you may also find my earlier post on spreadsheets as a common programming tool a useful foundation before diving into statistical methods.
Why SPC Matters in Process Monitoring
Semiconductor manufacturing processes require tight control over critical parameters to ensure consistent device performance, yield, and reliability. Across processes such as deposition, etch, lithography, and thermal treatments, even small variations in factors like temperature, pressure, gas flow, or power can lead to measurable shifts in material properties and device characteristics. Statistical Process Control (SPC) provides a real-time method for detecting process drift early, enabling engineers to identify and correct deviations before they result in out-of-specification wafers or reduced manufacturing yield.
Key SPC concepts for process monitoring include:
- X-bar chart: Tracks the average value of a critical process parameter (such as film thickness, critical dimension, or electrical performance) across subgroups, such as wafers or lots, over time to monitor process stability.
- R chart (range chart): Tracks the variability within each subgroup, helping identify changes in process consistency, uniformity, or equipment performance.
- Control limits: Statistically derived boundaries (typically ±3 standard deviations from the process mean) that define the expected range of normal process variation, distinct from engineering specification limits.
- Run rules: Additional statistical tests, such as multiple consecutive points above or below the center line, used to detect subtle trends, shifts, or non-random patterns before they become significant process issues.
Generating Synthetic Thin Film Thickness Data
For this walkthrough, we simulate thickness measurements from a hypothetical LPCVD silicon nitride process. This is a synthetic dataset created for illustrative purposes: we model 25 subgroups (batches) with 5 wafers each, a target thickness of approximately 2000 angstroms, and an intentional drift introduced in the final batches.
The synthetic data uses a mean of 2000 angstroms with a standard deviation of 15 angstroms, with a +5 angstrom per batch drift added in the last 5 batches. This mimics a real-world scenario where a chamber component such as a heating element begins to degrade, causing a gradual upward shift in deposited thickness.
We deliberately avoid claiming specific crystallographic orientations or substrate materials for this simulated data. The process environment is modeled as a generic polycrystalline thin film on a crystalline substrate, using peak intensity parameters consistent with typical nitride film characterization.
Building X-bar and R Charts in R
The R code below uses the qcc package, one of the most widely used libraries for SPC analysis. If you do not have it installed, run install.packages("qcc") first.
# Load package
library(qcc)
# Generate synthetic thickness data (angstroms)
set.seed(42)
n_batches <- 25
n_wafers <- 5
thickness <- matrix(nrow = n_batches, ncol = n_wafers)
for (i in 1:n_batches) {
drift <- ifelse(i > 20, (i - 20) * 5, 0)
thickness[i, ] <- round(rnorm(n_wafers, mean = 2000 + drift, sd = 15), 1)
}
# Create qcc objects
batch_thickness <- as.data.frame(thickness)
# X-bar chart
xbar_chart <- qcc(batch_thickness, type = "xbar",
title = "X-bar Chart: LPCVD Film Thickness",
xlab = "Batch Number", ylab = "Mean Thickness (A)")
# R chart
r_chart <- qcc(batch_thickness, type = "R",
title = "R Chart: LPCVD Film Thickness Range",
xlab = "Batch Number", ylab = "Range (A)")
Interpreting the Control Charts
When you run this code, the X-bar chart (shown below) will show all points within the upper and lower control limits for the first 20 batches. This indicates a stable, in-control process — exactly what a process engineer wants to see during routine production.
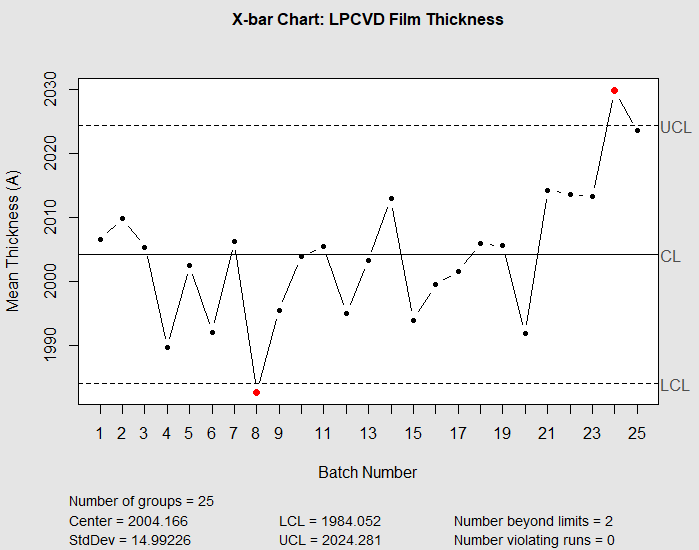
Beginning around batch 21, the mean thickness will begin to climb above the center line. By batch 23, one or more points may fall above the upper control limit (UCL), signaling that the process has shifted. The R chart should remain stable throughout, indicating that the within-batch uniformity (wafer-to-wafer variation) has not changed — the problem is a shift in the mean, not an increase in variability.
Applying Run Rules for Earlier Detection
Standard control limits alone may not detect gradual drifts quickly enough. Run rules add sensitivity:
- Rule 1: One point beyond the 3-sigma control limit.
- Rule 2: Seven consecutive points on the same side of the center line.
- Rule 3: Two out of three consecutive points beyond the 2-sigma warning limit.
In our simulated data, the 7-point run rule (Rule 2) would flag the drift as early as batch 22, before any individual measurement exceeds the control limits. This is the practical value of SPC: catching problems before they produce out-of-spec material.
R’s qcc package supports run rules via the rules argument. Adding rules = rulesets(c("rule1", "rule2", "rule3")) to the qcc() call will annotate violations directly on the chart.
Practical Applications for Process Engineers
SPC charts are not limited to thickness. They can be applied to any measurable property:
- Refractive index from ellipsometry measurements across a wafer batch.
- Film stress from wafer curvature measurements before and after deposition.
- Sheet resistance for conductive thin films measured by four-point probe.
- Uniformity calculated as (max – min) / (2 * mean) within a wafer.
The same R code structure shown above works for any of these variables. Simply replace the thickness data with your measurement values and the control chart logic remains identical.
Beyond Basic SPC: What Comes Next
Once you have SPC charts running, the next step is often process capability analysis (Cpk), which compares the natural process variation to the specification limits. A Cpk value below 1.33 typically indicates that the process needs improvement. We will cover capability analysis in a future post.
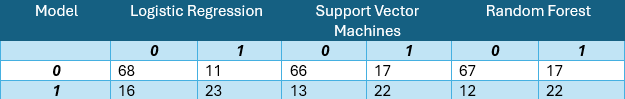
For readers interested in comparing multiple analytical approaches, our post on comparing logistic regression, SVMs, and random forests for classification demonstrates the kind of rigorous model comparison that complements SPC methodology in a broader data science toolkit.
Conclusion
SPC charts provide a straightforward, statistically grounded method for monitoring thin film deposition processes in real time. With just a few lines of R code using the qcc package, process engineers can detect drifts in mean thickness, identify changes in wafer-to-wafer variability, and trigger preventative maintenance before scrap material is produced. This blog uses a synthetic dataset to illustrate the workflow, but the same approach applies directly to real production data.
The combination of X-bar charts, R charts, and run rules gives process engineers a practical early warning system. In future posts, we will extend this framework to multivariate SPC (Hotelling’s T-squared) and explore how control charts integrate with broader statistical methods such as Design of Experiments (DOE), which we will cover in a subsequent blog.